Kai W. Wucherpfennig, M.D., Ph.D.
Professor and Co-Chair, Department of Cancer Immunology & AIDS

My laboratory works on the development of novel immunotherapies for the treatment of cancer.
Innovative Research in T cell Biology and Cancer Immunology:
T cells are key players in the immune system with the ability to detect and eliminate infected or transformed cells. We study the molecular mechanisms controlling antigen presentation to T cells, antigen recognition by the T cell receptor as well as activating and inhibitory signaling. We are particularly interested in T cell responses to self-antigens, which is beneficial in anti-tumor immunity yet detrimental in autoimmune diseases. Our projects aim to provide definitive mechanistic insights to key questions in the field. We use advanced tools to address these questions, ranging from in vivo studies to biochemistry, imaging and structural biology.
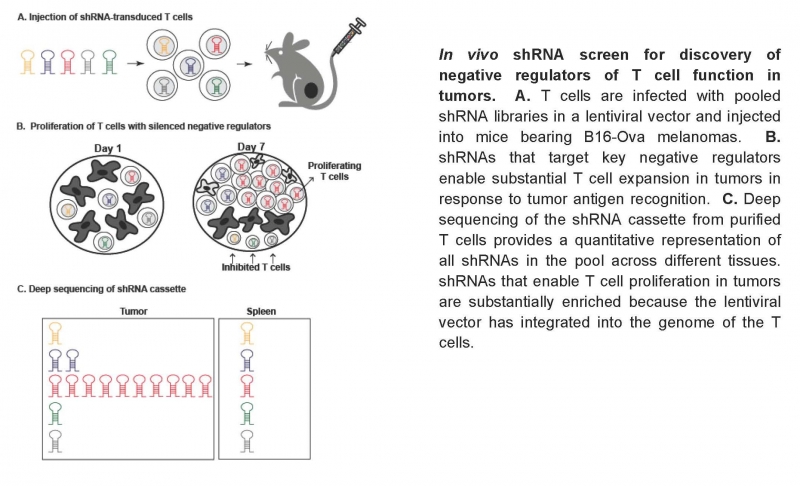
Recent work has shown that T cells play a central role in controlling tumor growth. Clinical trials have demonstrated that antibodies targeting two inhibitory receptors on the surface of T cells – CTLA-4 and PD-1 – can induce durable responses in a subset of cancer patients despite metastatic spread. We have recently developed a novel approach for in vivo discovery of genes that inhibit T cell function in the tumor microenvironment. In a normal immune response, T cells proliferate extensively following antigen recognition, but this physiological response is severely blunted by multiple immunosuppressive mechanisms in tumor microenvironments. We reasoned that shRNAs targeting critical negative regulators in T cells can restore their proliferative response to tumor antigen recognition.
We infect CD8 T cells with pool shRNA libraries and transfer these T cells into tumor-bearing mice. One week later, T cells are isolated to quantify the representation of shRNAs in tumors and control tissues by deep sequencing. We have discovered a large number of genes that inhibit T cell function in the tumor microenvironment. We now investigate the molecular mechanisms by which these genes constrain T cell function in tumors and use these insights to develop novel T cell-based therapeutic approaches.
Isolation of Human Antibodies that Enhance T cell mediated Tumor Immunity
Patients responding to a cancer immunotherapy developed by Dr. Glenn Dranoff at the Dana-Farber develop a robust antibody response. Some of these antibodies are directed against cell surface and secreted proteins that suppress T cell mediated anti-tumor immune responses.
We have developed a novel approach for isolation of circulating memory B cells from patients specific for such antigens. We find that fluorescent tetramers of the antigen of interest brightly label the relevant memory B cells. Single cell PCR of sorted B cells and sequencing of the immunoglobulin genes enables expression of recombinant antibodies for mechanistic studies. We first implemented this approach using tetanus toxoid as a human model antigen and found that the recombinant antibodies bound the antigen with high affinity. We are now using this approach to isolate human antibodies from patients responding to cancer immunotherapies, which is providing fascinating insights into mechanisms of immune protection against cancer. We believe that this is an exciting approach for the isolation of therapeutic antibodies.
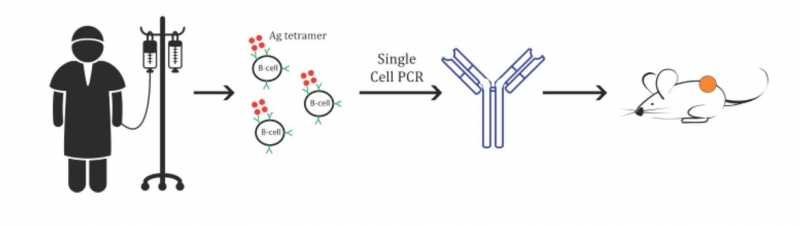 Isolation of human antibodies from cancer patients. Patients who respond to a cancer immunotherapy are tested for production of antibodies to tumor surface and secreted proteins. B cells that produce interesting antibodies are isolated with antigen tetramers. The therapeutic activity of recombinant antibodies is tested in a humanized mouse model.
Isolation of human antibodies from cancer patients. Patients who respond to a cancer immunotherapy are tested for production of antibodies to tumor surface and secreted proteins. B cells that produce interesting antibodies are isolated with antigen tetramers. The therapeutic activity of recombinant antibodies is tested in a humanized mouse model.
Molecular Mechanisms of T cell Recognition and Signaling
We are also studying key steps controlling T cell function. Of particular interest are the early events responsible for T cell receptor (TCR) activation following peptide antigen recognition. Using a novel live cell imaging approach, we showed that the CD3e cytoplasmic domain of the TCR binds to the inner leaflet of the plasma membrane in live T cells. Membrane binding was visualized by FRET between a C-terminal fluorescent protein and a membrane fluorophore. In collaboration with Dr. James Chou, we determined the NMR structure of the lipid-bound state of the CD3e cytoplasmic domain. The structure showed that the two key tyrosine residues partition into the hydrophobic core of the lipid bilayer. Sequestration of key tyrosine side chains into the lipid bilayer represents a novel mechanism for control of receptor activation.
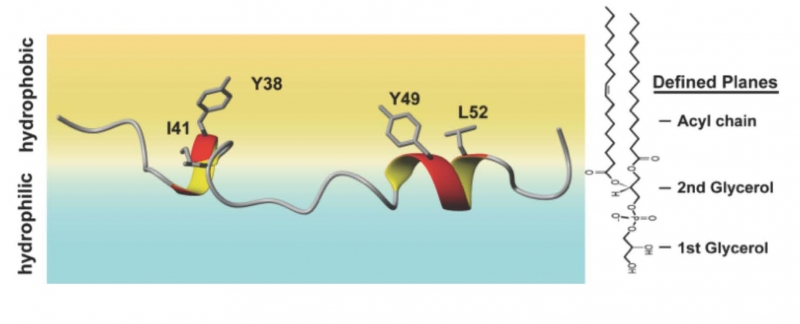 NMR structure of the CD3e ITAM in its membrane-bound state. The two tyrosine residues (Y38 and Y49) equilibrate into the hydrophobic acyl chain region (yellow) of the membrane.
NMR structure of the CD3e ITAM in its membrane-bound state. The two tyrosine residues (Y38 and Y49) equilibrate into the hydrophobic acyl chain region (yellow) of the membrane.
We have also shown that TCR triggering by peptide-MHC complexes induces dissociation of the CD3ε cytoplasmic domain from the plasma membrane. Electrostatic interactions between basic CD3e residues and acidic phospholipids are required for membrane binding. We observed rapid changes in the lipid composition of TCR microclusters in which signaling is initiated. These changes even occur when the proximal kinase in the activation cascade (Lck) is inhibited. Local changes in the lipid composition of TCR microclusters thus render the CD3ε cytoplasmic domain accessible during early stages of T cell activation.
We are also investigating other key regulatory events in T cell function. These studies are informed by the in vivo shRNA screen described above which has identified a large number of genes that control T cell function in the tumor microenvironment.
Molecular Mechanisms of Antigen Presentation of T cells
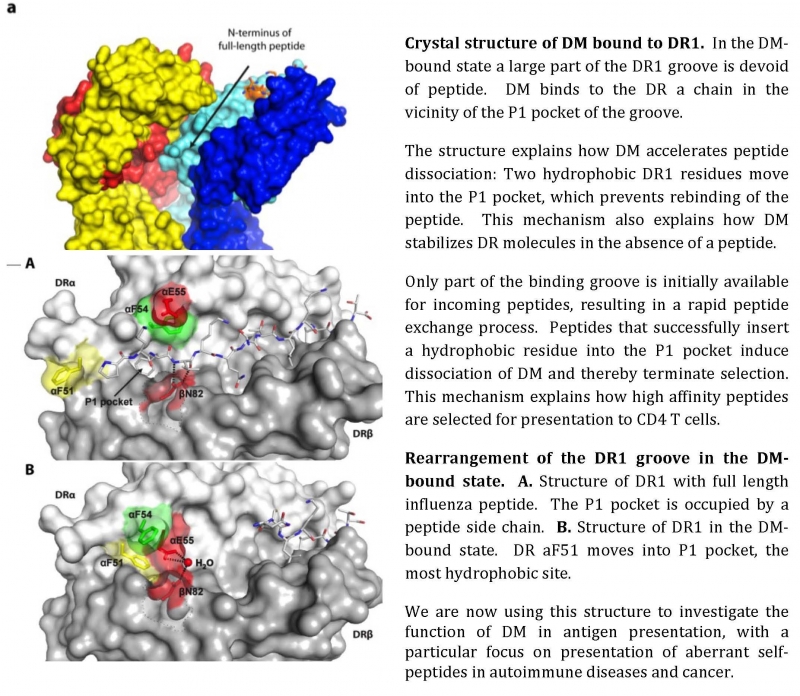
The repertoire of peptides bound to MHC proteins has a decisive impact on T cell mediated immunity. We study the biochemical mechanisms that control peptide selection in intracellular compartments. Of particular interest is HLA-DM (DM) which acts as an enzyme that accelerates selection of the highest-affinity peptides presented by MHC class II molecules. We have determined key biochemical steps in this interaction which allowed us to crystallize this important complex.
Selected Publications:
Zhou, P., Shaffer, D. R., Alvarez Arias, D. A., Nakazaki, Y., Pos, W., Torres, A. J., Cremasco, V., Dougan, S. K., Cowley, G. S., Elpek, K., Brogdon, J., Lamb, J., Turley, S. J., Ploegh, H. L., Root, D. E., Love, J. C., Dranoff, G., Hacohen, N., Cantor, H., and Wucherpfennig, K. W. (2014) In vivo discovery of immunotherapy targets in the tumour microenvironment, Nature 506, 52-57.
Pos, W., Sethi, D. K., Call, M. J., Schulze, M. S., Anders, A. K., Pyrdol, J., and Wucherpfennig, K. W. (2012) Crystal structure of the HLA-DM-HLA-DR1 complex defines mechanisms for rapid peptide selection, Cell 151, 1557-1568.
Anders, A. K., Call, M. J., Schulze, M. S., Fowler, K. D., Schubert, D. A., Seth, N. P., Sundberg, E. J., and Wucherpfennig, K. W. (2011) HLA-DM captures partially empty HLA-DR molecules for catalyzed removal of peptide, Nat Immunol 12, 54-61.
Xu, C., Gagnon, E., Call, M. E., Schnell, J. R., Schwieters, C. D., Carman, C. V., Chou, J. J., and Wucherpfennig, K. W. (2008) Regulation of T cell receptor activation by dynamic membrane binding of the CD3epsilon cytoplasmic tyrosine-based motif, Cell 135, 702-713.
Call, M. E., Pyrdol, J., Wiedmann, M., and Wucherpfennig, K. W. (2002) The organizing principle in the formation of the T cell receptor-CD3 complex, Cell 111, 967-979.
Contact Information
Smith Building, Room 736
450 Brookline Avenue,
Boston, MA 02115
